Why Does the Molecular Weight of My Protein Differ From the Theoretically Expected Weight? | Technology Networks

SAXSMoW 2.0: Online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale - Piiadov - 2019 - Protein Science - Wiley Online Library

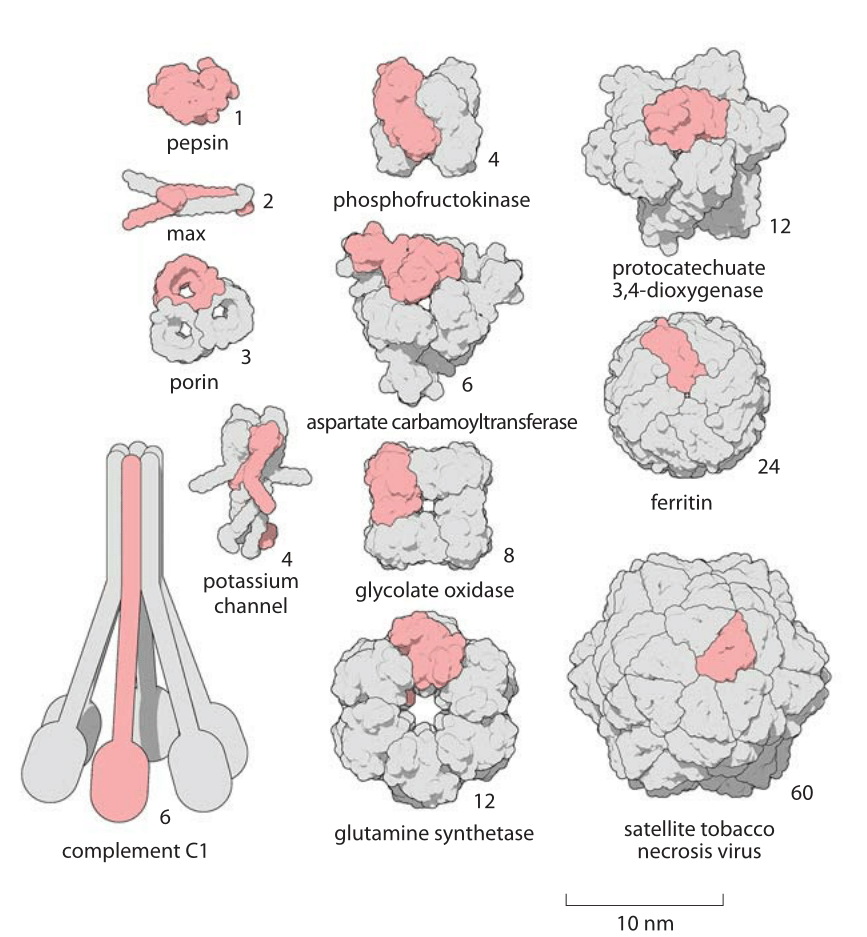
PDF) Size and Shape of Protein Molecules at the Nanometer Level Determined by Sedimentation, Gel Filtration, and Electron Microscopy
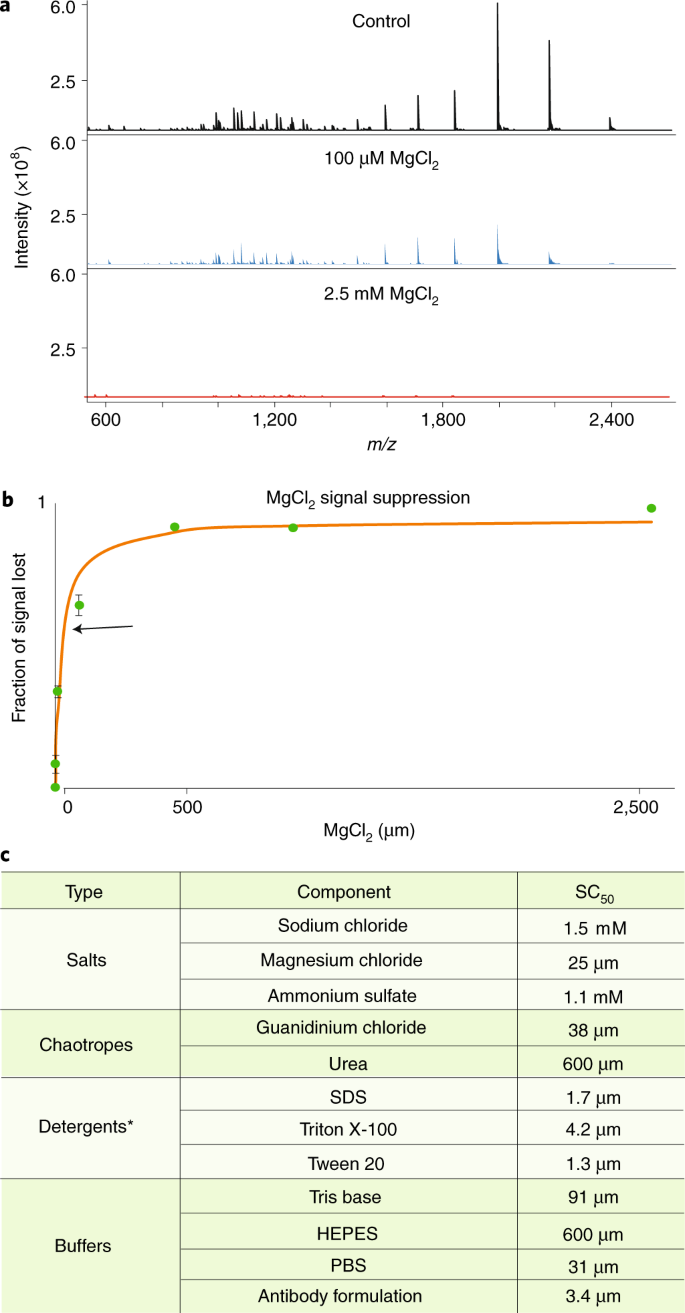
Best practices and benchmarks for intact protein analysis for top-down mass spectrometry | Nature Methods





/proteins1-4ac3f7a09ef84a879fdf30d4dc74056c.jpg)